MigenPro Package
Migenpro - Microbial Genome Prospecting
The increasing availability of microbial genomic data and recent development of machine learning and AI methods create a unique opportunity to establish associations between genetic information and phenotypes.
Research Overview
Migenpro serves as a framework for the generation of machine learning models that predict microbial traits from genome sequences. Microbial whole genome sequences have been consistently annotated and genomic features were stored in a semantic framework using the Genome Biology Ontology Language (GBOL).
Phenotype data corresponding to the available genome sequences was retrieved from BacDive and the associations between phenotype and genotype were used to train machine learning models.
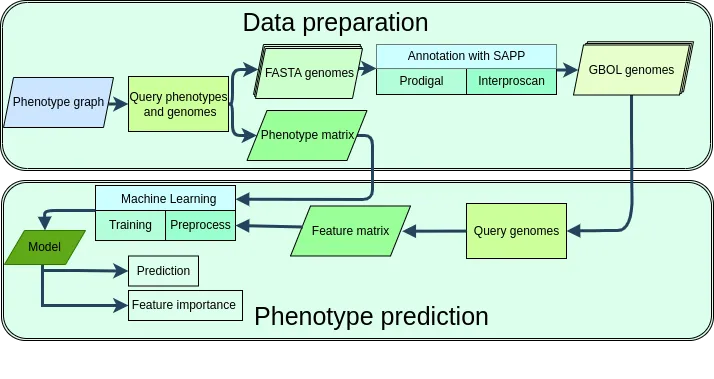
Key Results
- Successful Predictions: Our approach successfully predicts traits such as motility, Gram stain, optimal growth temperature range, and sporulation capabilities
- Robust Validation: Five-fold cross-validation demonstrated consistent model performance without overfitting
- Comparative Analysis: Comparable accuracy to previously published models with modest variations attributed to dataset differences
- Feature Analysis: Classification models can be explored using feature importance to identify biologically relevant genomic features
Impact
Migenpro provides an interoperable framework to predict phenotypes from genomic data, making bioprospecting more efficient for pharmaceuticals, agriculture, and environmental applications.